Structured Uncertainty
This page presents our ongoing work on modelling structured uncertainty for computer vision tasks. We introduce our Structure Uncertainty Prediction Network (SUPN) model published at CVPR 2018 working with Era Dorta and Lourdes Agapito.
Structured Uncertainty Prediction Networks
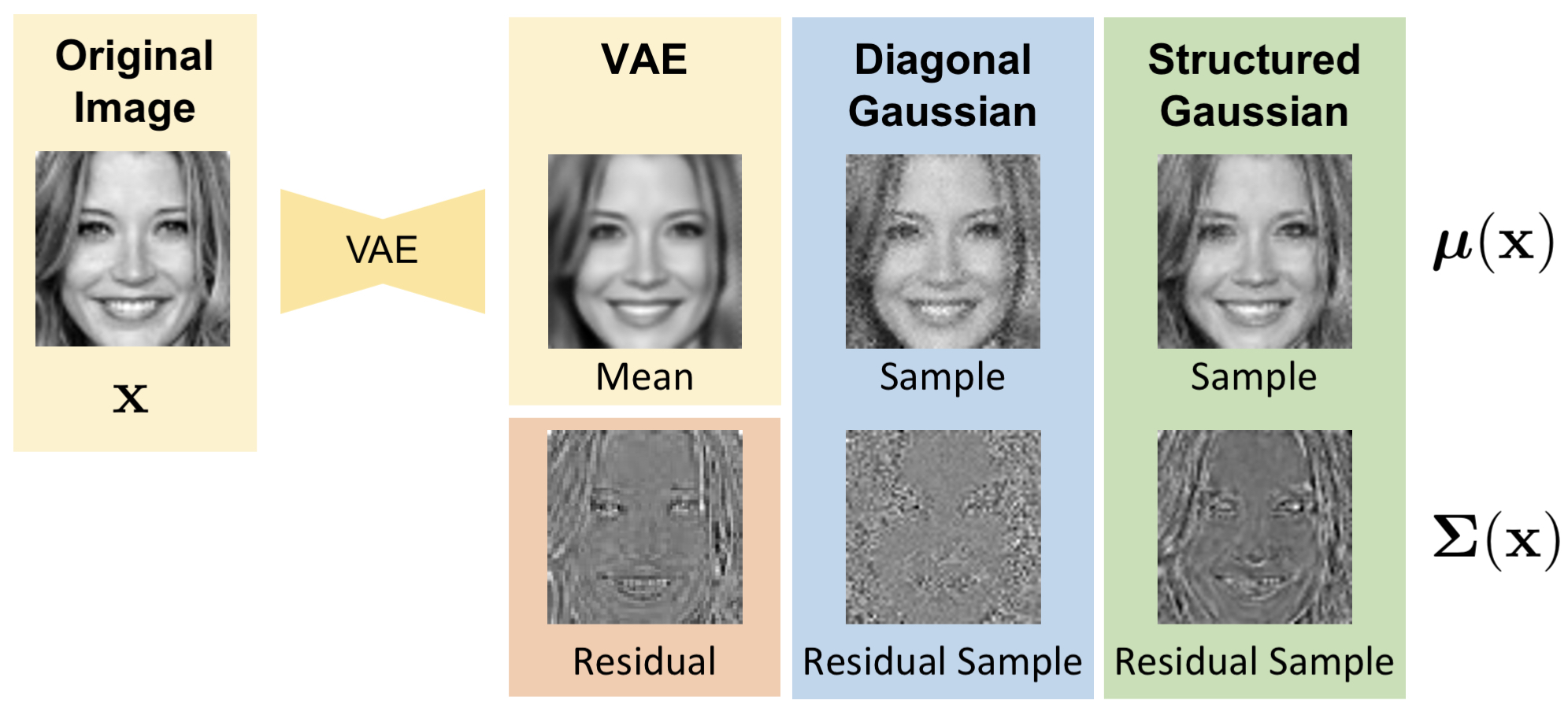
The figure provides an overview of our Structured Uncertainty Prediction Networks (SUPN) model. Consider a Variational Auto-Encoder (VAE) as a deep generative model. The reconstruction obtained from a VAE is often overly smooth due to the limitations of the architecture (e.g. latent space dimensionality) and the loss function used (e.g. Mean Squared Error or L2). We can consider the loss function as the log of the likelihood function over the observed data, thus the VAE predicts a distribution over the observed data. An L2 loss is the equivalent of a spherical Gaussian distribution with a single variance across all pixels.
To improve quality, it is common to estimate a diagonal distribution where we predict a different variance for each pixel. This is shown in the middle column. If we look at the residual between the mean VAE reconstruction and the original image (orange box) we can see that structure remains in the residual. The diagonal model fails to capture this structure; as a result, the error is estimated independently per pixel resulting in noisy samples that fail to reflect the original image statistics. In contrast, the SUPN model captures the structure in the error such that samples drawn are found to match the statistics of the residual and thus the original input.
Please see below for a section of a talk given at the BMVA workshop on generative models that discusses the SUPN model (PDF of slides):
Learning Structured Gaussians to Approximate Deep Ensembles
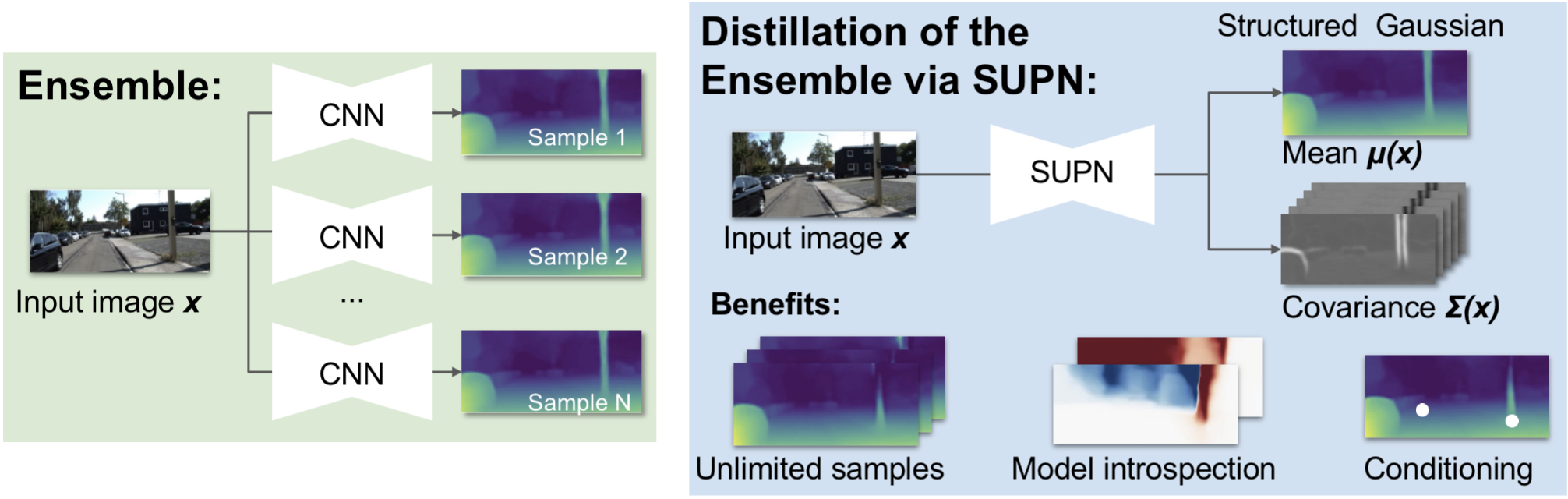
This work builds on our SUPN model to provide a closed-form approximator for the output of probabilistic deep ensembles used for dense image prediction tasks. Similarly to distillation approaches, the single network is trained to maximise the probability of samples from pre-trained probabilistic models, in the paper we use a fixed ensemble of networks. Once trained, this compact representation can be used to draw efficiently spatially correlated samples from the approximated output distribution. Importantly, this approach captures the uncertainty and structured correlations in the predictions explicitly in a formal distribution, rather than implicitly through sampling alone. This allows direct introspection of the model, enabling visualisation of the learned structure. Moreover, this formulation provides two further benefits: estimation of a sample probability, and the introduction of arbitrary spatial conditioning at test time. We demonstrate the merits of our approach on monocular depth estimation and show that the advantages of our approach are obtained with comparable quantitative performance.
Publications
Teo Deveney, Ivor Simpson and Neill D. F. Campbell,
Uncertainty for Safe Utilization of Machine Learning in Medical Imaging (UNSURE), LNCS, 2025
[pdf] [supplemental] [DOI: 10.1007/978-3-032-06593-3_22]
Margaret Duff, Ivor Simpson, Matthias J. Ehrhardt and Neill D. F. Campbell,
IoP Physics in Medicine and Biology, vol. 68, no. 16, 2023
[pdf] [arXiv link] [DOI: 10.1088/1361-6560/ace49a]
Ivor Simpson, Sara Vicente and Neill D. F. Campbell,
IEEE Conf. on Computer Vision and Pattern Recognition (CVPR), 2022
[pdf] [supplemental]
Era Dorta Perez, Sara Vicente, Lourdes Agapito, Neill D. F. Campbell and Ivor Simpson,
IEEE Conf. on Computer Vision and Pattern Recognition (CVPR), 2018
[pdf] [supplemental] [code]
Acknowledgements
This work has been supported by EPSRC CDE (EP/L016540/1) and CAMERA (EP/M023281/1) grants as well as the Royal Society.
